 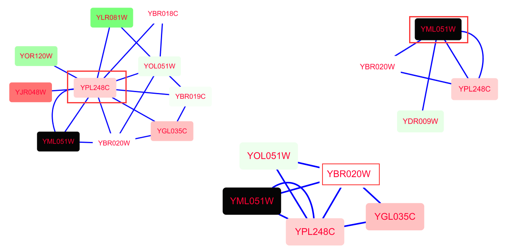
Gustavsen, Shraddha Pai, Ruth Isserlin, Barry Demchak, Alexander R. RC圓: Network biology using Cytoscape from within R Moreover, authors are now able to submit Study Protocols and Method Articles involving the broader Cytoscape ecosystem.Īs we look forward to the evolution of the Cytoscape Gateway, here’s a look at just some of the content that has helped shape this hub over the past seven years. In addition to the existing Software Tool articles that highlight Java-based Cytoscape Apps, the Gateway will also include technical extensions written using R or Python. To reflect the development of the Cytoscape platform over this period, we have updated the Gateway’s scope to encompass a greater variety of article topics. The Cytoscape Gateway has published many articles with tens of thousands of views and downloads between them. The Cytoscape Gateway is a tailored publication hub for new developments in the Cytoscape landscape. Significantly, Cytoscape also has non-biological applications. However, many also use the platform to interrogate the entire range from atomic and residue interactions in protein structures to chromosomal, cellular, organismal, and social networks. Researchers mainly use Cytoscape for molecular networks and pathways in biology. What is the Cytoscape Gateway?Ĭytoscape is an open-source software platform for network visualization, data integration, and network analysis. Here, Tom Sinden (Publishing Executive, F1000) shares new details about the Gateway’s recently expanded scope and looks back on some of the most important research to date. The Cytoscape Gateway has been a staple of the F1000Research bioinformatic landscape since launching in 2014.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
